NTHU Chemists Unveil the Nitrite-Reducing Machinery That Helps Staphylococcus aureus Survive (Dept. Chemistry / Prof. Yun-Wei Chiang)
Department: Department of Chemistry PI's Name: Yun-Wei Chiang
Journal: Structure and Nitrite Reductase Activity of the Di-iron Protein ScdA in Staphylococcus aureus, Journal of the American Chemical Society, 147 (2025) 31558-31569. https://doi.org/10.1021/jacs.5c05573
Title: NTHU Chemists Unveil the Nitrite-Reducing Machinery That Helps Staphylococcus aureus Survive
Abstract: Bacterial pathogens survive within the human host by counteracting intense chemical stresses generated by the immune system. Among these, reactive nitrogen species (RNS) represent a major antimicrobial strategy, yet the molecular mechanisms that enable pathogens to manage such stress remain incompletely understood. This study resolves a long-standing question surrounding ScdA, an enigmatic di-iron protein in Staphylococcus aureus, by revealing both its three-dimensional structure and its unexpected enzymatic function.
The central discovery is that ScdA is not merely a passive repair protein, as previously assumed, but a nitrite reductase that actively generates nitric oxide (NO). NO is a potent, double-edged molecule: while it is toxic at high concentrations, it also functions as a signaling and regulatory agent at controlled levels. Demonstrating that S. aureus encodes an enzyme capable of producing NO reframes our understanding of bacterial nitrosative stress responses and highlights a previously unrecognized strategy by which pathogens may manipulate host-derived chemistry.
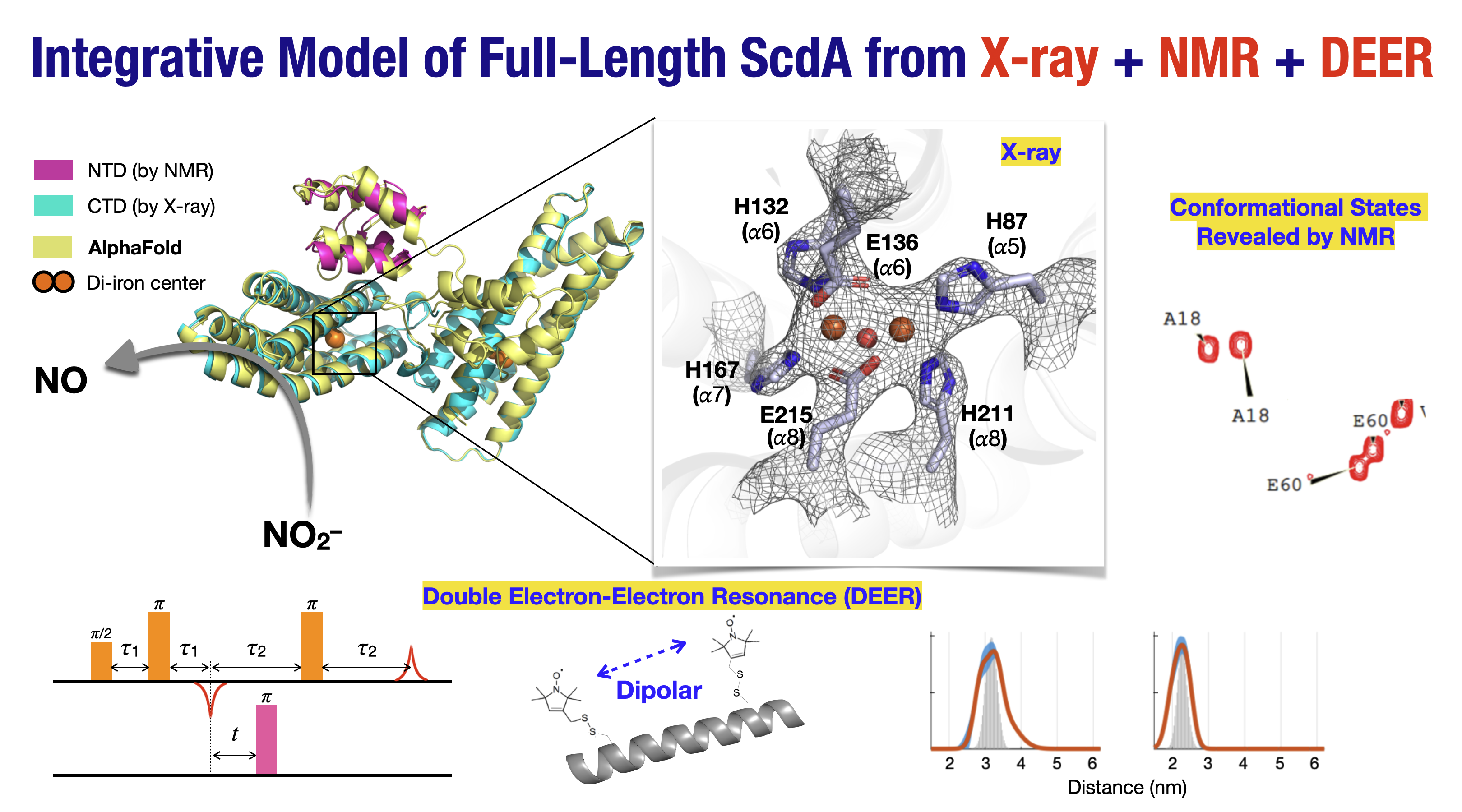
A major academic contribution of this work lies in its integrative structural biology approach. No single technique could resolve ScdA alone. X-ray crystallography defined the di-iron catalytic core, solution NMR elucidated the flexible N-terminal domain, AlphaFold modeling provided a full-length structural hypothesis, and pulsed EPR/DEER spectroscopy experimentally validated the dimeric architecture in solution. This represents one of the most comprehensive demonstrations to date of how AI-based structure prediction can be rigorously constrained and verified by advanced spectroscopy, establishing a broadly applicable methodological blueprint for studying dynamic, multi-domain proteins or even protein complexes.
Functionally, the study links structure to catalysis. Mutational analysis shows that dimerization and redox-sensitive cysteine chemistry tune enzymatic efficiency, revealing how oligomeric state controls substrate binding and turnover. Importantly, cellular assays demonstrate that ScdA-generated NO is biologically consequential, impairing bacterial growth when misregulated.
From a biomedical perspective, these findings identify NO homeostasis as a vulnerability in antibiotic-resistant pathogens, including MRSA. By uncovering how a stress-response protein can both protect and endanger the bacterium, this work opens new avenues for antimicrobial strategies that exploit redox imbalance rather than traditional antibiotic targets.